Enzymes have been instrumental in driving the evolution of Life on Earth and at Entropix, we believe they hold the key to solving many of our post-industrial challenges. Using the power of Directed Evolution, we are paving the way for enzymes to replace many of the conventional steps used in manufacturing. We are targeting approaches that have historically been associated with excessive energy demands and downstream pollution. Here we introduce the term the 3Es as a shorthand for the fundamental science underpinning our programmes at Entropix.
Enzymes Evolution and Entropy
All independent life forms depend on enzymes to orchestrate growth, reproduction, maintenance and death. And all enzymes are encoded by genes. The genome of a simple microbe like E.coli, encodes around 2000 different enzymes, and typically, the more sophisticated the organism, the more enzymes there are. In addition, throughout evolution, as species become more sophisticated, they have developed one or more layers of control over the intrinsic activity of their enzymes.
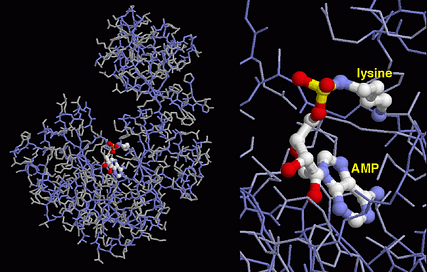
Although the information required to express enzymes takes the form of DNA, enzyme molecules are made up of hundreds of amino acids linked via a peptide bond to form a three-dimensional polymer. A specific sequence of bases (usually written in the single letter form as GATC) provides the information required to synthesise a particular chain of amino acids, which is either referred to as a protein or polypeptide. The enzyme molecule used in Covid PCR tests, Reverse Transcriptase, is shown on the right, with its substrate shown in light grey. This flow of information in living systems, from the gene to the protein is often described by the phrase


DNA makes RNA makes Protein
While double strands of DNA store the information needed to programme the expression of a cell’s enzymes, several forms of RNA are involved in the decoding process. Three major RNAs are required for translating the chemistry of nucleic acids into the chemistry of amino acids: messenger RNA (mRNA), ribosomal RNA (rRNA) and transfer RNA (tRNA). Through a highly controlled process called transcription, mRNA carries a copy of the information encoded by the gene to the ribosome. The ribosome, shown right is a supramolecular assembly of rRNA and a family of specialised proteins, that brings together mRNA and tRNA molecules specifying each amino acid. Ribosomes ensures that the information encoded in the genome is translated into the chemistry of amino acids, in an energy consuming process.
While the vast majority of enzymes are proteins, a small number are made from RNA: these are referred to as ribozymes. In fact, exploring the intimate interplay between nucleic acids and proteins has provided one of the richest areas of Scientific investigation for chemists and biologists alike.
Whether enzymes are derived from chains of amino acids or nucleotides, they have diverged and evolved to speed up a wide range of specific chemical reactions from protein degradation to DNA synthesis. Enzymes are the catalysts of life, but unlike the transition metals that catalyse many industrial processes, enzymes are capable of operating under relatively mild chemical conditions. This is one of the reasons that we believe enzymes represent the future of manufacturing and recycling industries.
Enzyme Active Sites
Enzymes typically engage their substrates in an intimate network of interactions in a highly organised pocket called the active site (shown in the expanded view on the left). The architecture of the active site is a unique product of the specific sequence of amino acids of each enzyme. It is at the heart of the active site where an enzyme persuades its substrates to develop into products with an efficiency that keeps the cellular programme on schedule.
Despite the efforts of many biochemists, over many years, our understanding of the molecular basis of enzyme catalysis remains incomplete. While these studies have provided us with many tools for measuring the substrate selectivity and the rates of enzymatic reactions, our ability to design enzymes from first principles remains elusive.
In the absence of a comprehensive understanding of the relationship between enzyme structure and catalysis, we rely on methodologies provided by molecular biologists to generate enzymes with novel properties. The method of Directed Evolution combined with high throughout sample handling technologies are employed at Entropix, to compress evolutionary time from many years into several weeks. Starting from a specific gene sequence, a library of mutants are first generated using a proprietary enzyme, and from this pool of variants, one or more candidate enzymes are selected to meet a set of pre-defined performance criteria, such as altered temperature or pH tolerance.
Evolution Embraces Diversity
The scale of the diversity of Life on Earth is a reflection of the many ways in which a genome can code for a living organism. Biologists like Charles Darwin relied on comparative anatomy and the fossil record to construct evolutionary relationships between the species. Today, we can look for the similarities between species by comparing the sequences of their genes and the proteins they encode. One of the most striking features to emerge from such molecular comparisons is that all species contain core genes, many of which are enzymes. An example of a core gene would be one essential for replication of the genome itself.
There are similarities between the amino acid sequences of enzymes that catalyse metabolic reactions in microbes and mammals. In some extreme cases the genes from mammals can carry out the functions of genes in yeast. However, with evolutionary time, the similarities between the sequences of enzymes have often “drifted” and the similarity becomes less easy to recognise. This divergence arises for a number of reasons: one that we exploit in Directed Evolution is the tendency of genome replication and repair enzymes to make the occasional mistake, producing a level of genetic variation that underpins evolutionary change. Using an in-house replication enzyme capable of controlled error-prone gene amplification, we can focus and accelerate natural levels of genetic variation, on a single gene.
The relationship between amino acid sequences within a class of enzyme or protein can sometimes be strictly limited, in which case the enzymes are said to be highly conserved. In other instances, there seems to be a much more relaxed, or less conserved retention of sequences over evolutionary time. The extent and the location of variation or conservation of sequence can be considered and evaluated in the context of information entropy.
Entropy is a measure of disorder in the Universe
The laws of thermodynamics state that all matter tends towards a state of disorder, rather than order. In information terms, this phenomenon was famously exploited by Claude Shannon as a way of expressing aspects of information in communication networks. The application of Shannon entropy forms the basis of one of the key computational tools that guides our Directed Evolution programmes at Entropix.
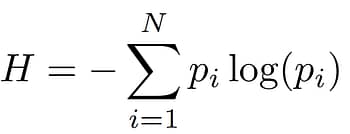
As discussed above, there is considerable evolutionary diversity between the sequences of enzymes carrying out specific functions: this is referred to as a high level of informational entropy. In contrast. some sequences are indispensable for function, and these are regions of low entropy. By systematically analysing the sequences of many variants generated both by Nature and through our in-house pipeline, we can reduce the time taken to search for the enzyme sequences that determine the new characteristics we desire.
The ultimate aim of all protein engineering is to rationally design enzymes with customised activity from first principles. This goal remains elusive and so in the meantime, Directed Evolution, informed by computational analysis, provides the best way forward for generating enzymes by learning from Nature and building on it.
In the next post, we shall discuss contemporary Bio-prospecting
