Introduction
Professor Simon Foster’s laboratory carries out pioneering research in the field of antibiotic resistance research. Uncovering the role of genomic elements in the resistance of Staphylococcus aureus to the penicillin class of antibiotics (MRSA) involves intensive screening of bacterial strains for alleles that promote resistance. Typically, genomic DNA is extracted from broth cultures and screened by PCR. Extraction of genomic DNA from gram positive strains is often more challenging than gram negatives, such as E. coli. However, using NADA™ technology, not only was the workflow successful, but the whole protocol reduced the timescale significantly. Dr Mariana Tinajero-Trejo, Research Associate at the University, found that NADA™ dramatically reduces processing times while maintaining exceptional DNA quality and yield, making it an ideal solution for high-throughput environments and time-sensitive diagnostics.

Experiment & Results
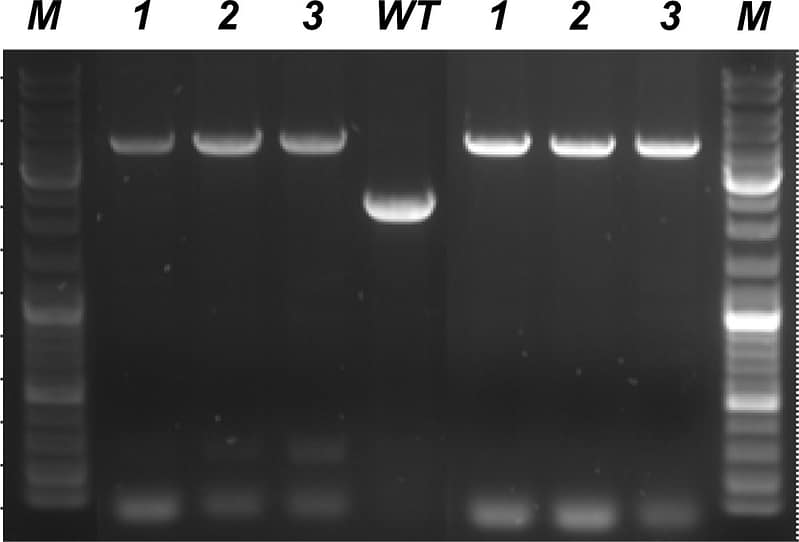
Extraction and purification of genomic DNA was carried out using NADA™ technology alongside a leading, commercial kit designed to extract genomic DNA from gram-positive bacteria (in this case MRSA SH1000 ΔsosA). The agarose gel below shows the results of a triplicate set of PCR amplifications, comparing genomic DNA purified by a leading commercial kit (left hand lanes 1-3), with those derived from NADA-ALGO™ purified DNA (right hand lanes 1-3). In all cases the major amplicons were of the correct molecular weight (lanes M are molecular weight standards). Importantly, as expected, the mutant alleles were longer than those derived from a wild-type strain (lane WT). In addition, the amplicon yield was higher in the NADA-ALGO™ samples (compare left and right hand sets of lanes) and no non-specific low molecular weight amplification was observed with NADA-ALGO™ purified templates. The NADA-ALGO™ protocol took 10 mins from cell harvesting and lysis: the competitor protocol took 60 mins, involving multiple centrifugations and spin-column bind-wash-elute steps.
Professor David Hornby
CSO and co-founder of Entropix
“This work highlights the potential of NADA™ to transform laboratory efficiency and reliability, offering a powerful tool for academic, clinical, and industrial labs alike. As the demand for fast, accurate genetic analysis continues to grow, NADA™ technology from Entropix stands at the forefront of next-generation DNA purification.”
Dr Mariana Tinajero-Trejo
Post Doctoral Research Fellow University of Sheffield
“The NADA™ process was so rapid and easy to use compared with the commercial kits, which require repeated sample washing steps. It’s also more sustainable with fewer single-use plastic tubes”
For further information, samples or trials, please contact enquiries@entropix.co.uk
Download a copy of the case study.
